suggests: คุณกำลังดูกระทู้
At least 30 outbreaks of Ebola virus disease (EVD) have been identified since the late 1970s, the most severe of which affected Guinea, Sierra Leone and Liberia from December 2013 to June 20161,2. Guinea experienced a new outbreak of EVD in 2021, which started in Gouéké—a town about 200 km away from the epicentre of the 2013–2016 outbreak. The probable index case was a 51-year-old nurse, an assistant of the hospital midwife in Gouéké. On 21 January 2021, she was admitted to hospital in Gouéké suffering from headache, asthenia, nausea, anorexia, vertigo and abdominal pain. She was diagnosed with malaria and salmonellosis and was released two days later. Feeling ill again once at home, she attended a private clinic in Nzérékoré (40 km away) and visited a traditional healer, but died three days later. In the week after her death, her husband—as well as other family members who attended her funeral—fell ill, and four of them died. They were reported as the first suspect cases by the national epidemic alert system on 11 February. On 12 February, blood was taken from two suspect cases admitted to hospital in Nzérékoré. On 13 February, both of these patients were confirmed to have EVD by the laboratory in Guéckédou, which used a commercial real-time polymerase chain reaction with reverse transcription (qRT–PCR) assay (RealStar Filovirus Screen Kit, Altona Diagnostics). On 13 February, the husband of the index case—who travelled more than 700 km from Gouéké to Conakry, the capital city of Guinea, for treatment—was admitted to the Centre de Traitement Epidémiologique (CTEpi) in Nongo, Ratoma Commune. He presented with fever, nausea, asthenia, abdominal pain and lumbar pain and was strongly suspected to have EVD. A blood sample was analysed on the same day and was found to be positive for Ebola Zaire (; EBOV) according to the GeneXpert molecular diagnostic platform (Xpert Ebola test, Cepheid) and by an in-house qRT–PCR assay. Laboratory confirmation of EVD in the three suspect cases led to the official declaration of the epidemic on 14 February. By 5 March, 14 confirmed cases and 4 probable cases of EVD had been identified, leading to 9 deaths—including 5 confirmed cases as reported by the Agence Nationale de la Sécurité Sanitaire (ANSS) of Guinea. After a period of 25 days without new cases, two new cases were reported around Nzérékoré on 1 and 3 April, and on 19 June 2021 the outbreak was declared to be over. In total, 16 confirmed cases were reported, among which 12 people died.
Genomic characterization of the virus that caused the 2021 epidemic of EVD in Guinea was of immediate importance to public health. First, because diagnostic tools, therapeutics and vaccines with proven effectiveness in recent EVD outbreaks—such as in Guinea (2013–2016) and in the Equateur and North-Kivu/Ituri provinces of the Democratic Republic of the Congo (DRC) (2018–2020)—have primarily been developed for EBOV3,4,5. Second, to identify whether the outbreak resulted from a new zoonotic transmission event or from the resurgence of a viral strain that had circulated in a previous EBOV outbreak: it is known that EBOV can persist in the bodily fluids of patients who have survived EVD and can be at the origin of new transmission chains6,7,8. Although the Xpert Ebola test was developed to detect only EBOV strains and the in-house qRT–PCR assay uses a probe that is specifically designed to detect EBOV9, additional confirmation by sequence analysis was sought by targeting a short fragment in the viral protein 35 region of the sample from the patient who was hospitalized in Conakry. The phylogenetic tree (Supplementary Fig. 1) underscores that this highly conserved region can discriminate between Ebola virus species, and analysis confirmed that the virus that caused the new outbreak was of the species . This confirmed that available vaccines and the vast majority of molecular-diagnostic tools and therapeutics could be immediately applied.
To gain further insight into the genomic make-up of the viruses causing this outbreak, 11 complete or near-complete (greater than 95% recovery) and 8 partial (greater than 65% recovery) genomic sequences from 12 of the 14 confirmed cases were obtained by 3 different laboratories using different next-generation sequencing technologies (Table 1). To facilitate the public-health response and the evaluation of existing medical countermeasures, sequencing results were made publicly available on 12 March through joint posting (https://virological.org/c/ebolavirus/guinea-2021/44). Blood and swab samples from 14 patients with confirmed EVD, sampled from 12 February to 4 March, were processed by the following methods: hybridization capture technology and sequencing on Illumina iSeq100, an amplicon-based protocol with EBOV-specific primer pools and sequencing on MinION (Oxford Nanopore Technologies (ONT)), and a hybrid-capture-based approach using a probe panel that included EBOV-specific targets followed by TruSeq exome enrichment, as previously described5. The data generated between the three groups were pooled and the sequence that had the highest quality was chosen for each patient. This enabled us to reconstruct 12 high-quality EBOV genomes that covered 82.9–99.9% of the reference genome (KR534588) (Table 1). The consensus EBOV sequences with the highest genome recovery (greater than 82.9%) from 12 different patients were used in further analyses.
Table 1 Patient and sample characteristics and sequencing results obtained by the laboratories involved in the study
Full size table
Maximum likelihood phylogenetic reconstruction places the 12 genomes from the 2021 outbreak of EVD in Guinea as a single cluster among the EBOV viruses that were responsible for the 2013–2016 outbreak in West Africa (Figs. 1, 2). The genomes from the 2021 outbreak share 10 substitutions (compared with KJ660346) that were accumulated during the 2013–2016 outbreak, including the A82V marker mutation for human adaptation in the glycoprotein that arose when the virus spread to Sierra Leone11,12. These patterns provide strong evidence of a direct link to human cases from the 2013–2016 outbreak rather than a new spillover from an animal reservoir. The 2021 lineage is nested within a clade that predominantly consists of genomes sampled from Guinea in 2014 (Fig. 2). The branch by which the 2021 cluster diverges from the previous outbreak exhibits only 12 substitutions, which is far fewer than would be expected from the evolution of EBOV during 6 years of sustained human-to-human transmission (Fig. 3). Using a local molecular-clock analysis, we estimate a 6.4-fold (95% highest posterior density (HPD) interval: 3.3-fold, 10.1-fold) lower rate along this branch. For comparison, we also estimate a 5.5-fold (1.6-fold, 10.8-fold) lower rate along the branch leading to the 2016 cases, which were linked to a patient who survived the disease and in whom the virus persisted for more than 500 days7,13. Rather than a constant long-term low evolutionary rate, some degree of latency or dormancy during persistent infection seems to be a more likely explanation for the low divergence of the genomes from the 2021 epidemic. We tested whether the 12 genomes from the 2021 epidemic, which were sampled over a time period of less than one month, contained sufficient temporal signal to estimate the time to most recent common ancestor (tMRCA) (Supplementary Fig. 2); however, we did not identify statistical support for sufficient divergence accumulation over this short timescale. We therefore calibrated our analysis using an evolutionary rate that reflects EBOV evolution under sustained human-to-human transmission (as estimated by the local molecular-clock analysis). This resulted in a tMRCA estimate of 22 January 2021 (95% HPD interval: 29 December 2020, 10 February 2021).
Fig. 1: Maximum likelihood phylogenetic reconstruction for 55 representative genomes from previous outbreaks of and 12 genomes from the 2021 outbreak in Guinea.
Most clades for single or multiple closely related outbreaks are collapsed and internal node support is proportional to the size of the internal node circles. The clades or tip circles are labelled with the locations and years of the outbreaks, and coloured according to the (first) year of detection.
Full size image
Fig. 2: Maximum likelihood phylogenetic reconstruction for 1,065 genomes sampled during the 2013–2016 West African outbreak and 12 genomes from the 2021 outbreak in Guinea.
A colour gradient (from purple to green for increasing divergence) is used to colour the tip circles. The 2021 genomes are shown with a larger circle in yellow.
Full size image
Fig. 3: Temporal divergence plot showing genetic divergence from the root over time.
This plot relates to the tree shown in Fig. 2. The regression is exclusively fitted to genomes sampled between 2014 and 2015. The same colours are used for the data points as in Fig. 2. The dashed yellow lines highlight how the 2021 data points deviate from the relationship between sampling time and sequence divergence. According to this relationship, about 95 substitutions (95% prediction interval: 88–101) are expected on the branch ancestral to the 2021 cluster, whereas only 12 are inferred on this branch.
Full size image
These results open up a new perspective on the relatively rare observation of EBOV re-emergence. It is assumed that all known filovirus outbreaks in humans are the result of independent zoonotic transmission events from bat reservoir species or from intermediate or amplifying hosts such as apes and duikers6. Here we clearly show that, even almost five years after the declaration of the end of an epidemic, new outbreaks could also be the result of transmission from humans who were infected during a previous epidemic. The viruses from the 2021 outbreak fall within the lineage of EBOV viruses obtained from humans during the 2014–2016 outbreak; as such, it is very unlikely that this new outbreak has an animal origin or is the result of a new cross-species transmission with the same lineage that remained latent in this natural host, which in that scenario would be at the basis of the West African cluster. The limited genomic divergence between 2014–2015 and 2021 is compatible with a slow long-term evolutionary rate. However, a relatively long phase of latency might be more likely than continuous slow replication. Independent of the mechanistic explanation, the virus most probably persisted at a low level in a human who had survived previous infection. Plausible scenarios of EBOV transmission to the index case include: sexual transmission by exposure to EBOV in semen from a male survivor; contact with body fluids from a survivor who had a relapse of symptomatic EVD (for example during healthcare—the index case was a healthcare worker); or relapse of EVD in the index case—although she was not known to have been infected previously, she could have had an asymptomatic or pauci-symptomatic EBOV infection during the previous outbreak. A detailed investigation of the family of the index case by anthropologists revealed that she was not known to have had EVD previously, nor were her husband or close relatives. However, among more distantly related family, 25 individuals had EVD during the previous outbreak. Only five survived, although the index case apparently had no recent contacts with this part of the family. Consultation of the hospital registers in Gouécké showed that all patients seen by the index case in January 2021 were in good health and were still in good health in March 2021. However, the index case also performed informal consultations outside the hospital environment, which could not be verified. An alternative scenario is that the nurse was not the actual index case, but was part of a small, unrecognized chain of human-to-human transmission in this area of Guinea. However, the diversity of the currently available genomes is limited, and molecular-clock analysis suggests a recent tMRCA, with a mean estimate close to the time that the nurse was first hospitalized and a 95% HPD boundary around the beginning of the year. This provides some reassurance that the outbreak was detected early.
The 2013–2016 outbreak in West Africa was the largest and most complex recent outbreak of EBOV, and involved more than 28,000 cases, 11,000 deaths and an estimated 17,000 survivors, mostly in Guinea, Liberia and Sierra Leone2. This large outbreak provided new information about the disease itself as well as about the medical, social and psychological implications for patients who survived the disease14,15,16. It was also possible to estimate, to some extent, the proportions of asymptomatic or pauci-symptomatic infections and to identify their role in specific unusual transmission chains17,18,19. Although the main route of human-to-human transmission of EBOV is direct contact with infected bodily fluids from symptomatic or deceased patients, some transmission chains in this outbreak were associated with viral persistence in semen3. Several studies demonstrated viral persistence in more than 50% of male survivors at 6 months after discharge from Ebola treatment units (ETU), and the maximum duration of persistence in semen has been reported to be up to 500–700 days after ETU discharge in a small number of male EVD survivors9,20,21,22. Transmission through other bodily fluids (such as breast milk and cervicovaginal fluids) is also suspected8,23,24,25. Furthermore, some immunological studies among survivors suggest a continuous or intermittent EBOV antigenic stimulation due to persistence of an EBOV reservoir in some survivors26,27, although this was not confirmed in another study28. Cases of relapse of EVD have also been sporadically reported and could be the origin of large transmission chains, as recently reported in the North-Kivu outbreak in DRC29. For example, the presence of EBOV RNA, 500 days after ETU discharge, in the breast milk of a woman who was not pregnant when she developed EVD has recently been reported. She attended the hospital owing to complications at 8 months of pregnancy, and a breast milk sample that was taken 1 month after delivery tested positive for EBOV RNA9. These examples illustrate that healthcare workers can be exposed to EBOV when taking care of patients who survived EVD but have an unrecognized relapse of their infection. The 2021 outbreak now highlights that viral persistence and reactivation is not limited to a two-year period, but can also occur on much longer timescales with late reactivation.
Active genomic surveillance has already shown the resurgence of previous strains in other outbreaks of the disease. For example, two EBOV variants circulated simultaneously within the same region during the recent 2020 outbreak in Equateur province, DRC30. Moreover, strains from the two consecutive outbreaks in Luebo, DRC, in 2007 and 2008, are also so closely related that it now seems difficult to exclude that the epidemic observed in 2008 was due to a resurgence event from patient who survived EVD in the 2007 outbreak31,32. However, the limited genomic sampling does not allow for a formal test of this hypothesis.
Although the majority of EVD outbreaks remained limited both in the number of cases and in geographic spread, the two largest outbreaks in West Africa (December 2013–June 2016) and in eastern DRC (August 2018–June 2020) infected thousands of individuals over wide geographic areas, leading to large numbers of EVD survivors. This means that the risk of resurgence is higher than ever before. Continued surveillance of EVD survivors is therefore warranted to monitor the reactivation and relapse of EVD infection and the potential presence of the virus in bodily fluids. This work and associated communications must be conducted with the utmost care for the wellbeing of EVD survivors. During the 2013–2016 outbreak in Guinea, patients who survived EVD had a mixed experience after discharge from ETUs. On the one hand, they were considered as heroes by non-governmental organizations and became living testimonies of a possible recovery33,34. On the other hand, they experienced different forms of stigmatization, such as rejection by family and friends, refusal of involvement in collective work, loss of jobs and housing, and sometimes self-isolation from social life and workplaces35. The human origin of the 2021 EVD outbreak, and the associated shift in our perception of EBOV emergence, call for careful attention to survivors of the disease. The concern that survivors will be stigmatized as a source of danger should be a matter of scrupulous attention36. This is especially true for the area of Gouécké, which is only 9 km away from Womey—a village that is emblematic of the violent reaction of the population towards the EVD response team during the 2013–2016 epidemic37.
Since the 2013–2016 EVD outbreak in Western Africa, genome sequencing has become a major component of the response to outbreaks10,38,39,40,41. The establishment of in-country sequencing and the building of capacity enabled a timely characterization of EBOV strains in the 2021 outbreak in Guinea. In addition to the importance of appropriate healthcare measures focused on survivors, the late resurgence of the virus also highlights the urgent need for further research into potent antiviral agents that can eradicate the latent virus reservoir in patients with EVD, and into efficient vaccines that provide long-term protection. In parallel, vaccination could also be considered to boost protective antibody responses in survivors of the disease27. The vaccination of populations in areas with previous EBOV outbreaks could also be promoted to prevent secondary cases.
Table of Contents
[NEW] Bizarre dermal armour suggests the first African ankylosaur | suggests – NATAVIGUIDES
-
1.
Vickaryous, M. K., Maryańska, T. & Weishampel, D. B. in (eds Weishampel, D. B. et al.) 363–392 (Univ. California Press, 2004).
-
2.
Thompson, R. S., Parish, J. C., Maidment, S. C. R. & Barrett, P. M. Phylogeny of the ankylosaurian dinosaurs. 10, 301–312 (2012).
-
3.
Arbour, V. M. & Currie, P. J. Systematics, phylogeny and palaeobiogeography of the ankylosaurid dinosaurs. 14, 385–444 (2016).
-
4.
Maidment, S. C. R., Raven, T. J., Ouarhache, D. & Barrett, P. M. North Africa’s first stegosaur: implications for Gondwanan thyreophoran dinosaur diversity. 77, 82–97 (2020).
-
5.
Galton, P. M. Lydekker, an ankylosaurian dinosaur from the Middle Jurassic of England. 1983, 141–155 (1983).
-
6.
Leahey, L., Molnar, R. E., Carpenter, K., Witmer, L. M. & Salisbury, S. W. Cranial osteology of the ankylosaurian dinosaur formerly known as sp. (Ornithischia: Thyreophora) from the Lower Cretaceous Allaru Mudstone of Richmond, Queensland, Australia. 3, e1475 (2015).
-
7.
Owen, R. Report on British fossil reptiles. 11, 60–204 (1842).
-
8.
Seeley, H. G. On the classification of the fossil animals commonly named Dinosauria. 43, 258–265 (1888).
-
9.
Osborn, H. F. Two Lower Cretaceous dinosaurs of Mongolia. 95, 1–10 (1923).
-
10.
de Buffrénil, V., Farlow, J. O. & de Ricqlès, A. Growth and function of plates: evidence from bone histology. 12, 459–473 (1986).
-
11.
Scheyer, T. M. & Sander, P. M. Histology of ankylosaur osteoderms: implications for systematics and function. 24, 874–893 (2004).
-
12.
Scheyer, T. M., Sander, P. M., Joyce, W. G., Bohme, W. & Witzel, U. A plywood structure in the shell of fossil and living soft-shelled turtles (Trionychidae) and its evolutionary implications. 7, 136–144 (2007).
-
13.
Berkovitz, B. & Shellis, R. P. (Academic Press, 2017).
-
14.
Schnetz, L., Pfaff, C. & Kriwet, J. Tooth development and histology patterns in lamniform sharks (Elasmobranchii, Lamniformes) revisited. 277, 1584–1598 (2016).
-
15.
Jambura, P. L. et al. Micro-computed tomography imaging reveals the development of a unique tooth mineralization pattern in mackerel sharks (Chondrichthyes, Lamniformes) in deep time. 9, 9652 (2019).
-
16.
Jambura, P. L. et al. Evolutionary trajectories of tooth histology patterns in modern sharks (Chondrichthyes, Elasmobranchii). 236, 753–771 (2020).
-
17.
Reid, R. E. H. Bone histology of the Cleveland-Lloyd dinosaurs and of dinosaurs in general, part 1: introduction to bone tissues. 41, 25–72 (1996).
-
18.
Main, R. P., de Ricqlès, A., Horner, J. R. & Padian, K. The evolution and function of thyreophoran dinosaur scutes: implications for plate function in stegosaurs. 31, 291–314 (2005).
-
19.
Dubansky, B. H. & Dubansky, B. D. Natural development of dermal ectopic bone in the American alligator () resembles heterotopic ossification disorders in humans. 301, 56–76 (2018).
-
20.
Kirby, A. et al. A comparative histological study of the osteoderms in the lizards (Squamata: Helodermatidae) and (Squamata: Varanidae). 236, 1035–1043 (2020).
-
21.
Shadwick, R. E., Russell, A. P. & Lauff, R. F. The structure and mechanical design of rhinoceros dermal armour. 337, 419–428 (1992).
-
22.
Hall, B. K. (Elsevier, 2015).
-
23.
Galton, P. M. & Upchurch, P. in (eds Weishampel, D. B. et al.) 343–362 (Univ. California Press, 2004).
-
24.
Barrett, P. M., Clarke, J. B., Brinkman, D. B., Chapman, S. D. & Ensom, P. C. Morphology, histology and identification of the ‘granicones’ from the Purbeck Limestone Formation (Lower Cretaceous: Berriasian) of Dorset, southern England. 23, 279–295 (2002).
-
25.
Burns, M. E. & Currie, P. J. External and internal structure of ankylosaur (Dinosauria, Ornithischia) osteoderms and their systematic relevance. 34, 835–851 (2014).
-
26.
Norman, D. B. (Dinosauria: Ornithischia) from the Early Jurassic of Dorset, England: biology and phylogenetic relationships. 191, 1–86 (2021).
-
27.
Scheyer, T. M., Syromyatnikova, E. V. & Danilov, I. G. Turtle shell bone and osteoderm histology of Mesozoic and Cenozoic stem-trionychian Adocidae and Nanhsiungchelyida (Crypodira: Adocusia) from Central Asia, Mongolia and North America. 20, 69–85 (2017).
-
28.
Cerda, I. A., Desojo, J. B. & Scheyer, T. M. Novel data on aetosaur (Archosauria: Pseudosuchia) osteoderm microanatomy and histology: palaeobiological implications. 61, 721–745 (2018).
-
29.
D’Emic, M. D., Wilson, J. A. & Chatterjee, S. The titanosaur (Dinosauria: Sauropoda) osteoderm record: review and first definitive specimen from India. 29, 165–177 (2009).
-
30.
Scheyer, T. M., Desojo, J. B. & Cerda, I. A. Bone histology of phytosaur, aetosaur, and other archosauriform osteoderms (Eureptilia, Archosauromorpha). 297, 240–260 (2014).
-
31.
Cerda, I. A., García, R. A., Powell, J. E. & Lopez, O. Morphology, microanatomy and histology of titanosaur (Dinosauria, Sauropoda) osteoderms from the Upper Cretaceous of Patagonia. 35, e905791 (2015).
-
32.
Stocker, M. R. & Butler, R. J. in (eds Nesbitt, S. J. et al.) 91–117 (Geological Society, London, 2013).
-
33.
Desojo, J. B., et al. in (eds Nesbitt, S. J. et al.) 203–239 (Geological Society, London, 2013).
-
34.
Gaffney, E. S. The comparative osteology of the Triassic turtle . 194, 1–263 (1990).
-
35.
Gorsack, E. & O’Connor, P. M. Time-calibrated models support congruency between Cretaceous continental rifting and titanosaurian evolutionary history. 12, 20151047 (2016).
-
36.
Maidment, S. C. R. Stegosauria: a historical review of the body fossil record and phylogenetic relationships. 103, 199–210 (2010).
-
37.
Reichel, M., Schultz, C. L. & Soares, M. B. A new traversodontid cynodont (Therapsida, Eucynodontia) from the Middle Triassic Santa Maria Formation of Rio Grande do Sul, Brazil. 52, 229–250 (2009).
-
38.
Scheyer, T. M. & Sues, H. D. Expanded dorsal ribs in the Late Triassic pseudosuchian reptile . 37, e1248768 (2017).
-
39.
Salgado, L. et al. A new primitive neornithischian dinosaur from the Jurassic of Patagonia with gut contents. 7, 42778 (2017).
-
40.
Molnar, R. E. & Wiffen, J. A Late Cretaceous polar dinosaur fauna from New Zealand. 15, 689–706 (1994).
-
41.
Coria, R. A. & Salgado, L. in (ed. Carpenter, K.) 159–168 (Indiana Univ. Press, 2001).
-
42.
Nath, T. T., Yadagiri, P. & Moitra, A. K. First record of armoured dinosaur from the Lower Jurassic Kota Formation, Pranhita-Godavari Valley, Andhra Pradesh. 59, 575–577 (2002).
-
43.
de Valais, S., Apesteguía, S. & Udrizar Sauthier, D. Neuvas evidencias de dinosaurios de la Formación Puerto Yeruá (Cretacico), Provincia de Entre Ríos, Argentina. 40, 631–635 (2003).
-
44.
Salgado, L. & Gasparini, Z. Reappraisal of an ankylosaurian dinosaur from the Upper Cretaceous of James Ross Island (Antarctica). 28, 119–135 (2006).
-
45.
Barrett, P. M. et al. Ankylosaurian dinosaur remains from the Lower Cretaceous of southeastern Australia. 34, 205–217 (2010).
-
46.
Leahey, L. G. & Salisbury, S. W. First evidence of ankylosaurian dinosaurs (Ornithischia: Thyreophora) from the mid-Cretaceous (late Albian–Cenomanian) Winton Formation of Queensland, Australia. 37, 249–257 (2013).
-
47.
Galton, P. M. Earliest record of an ankylosaurian dinosaur (Ornithischia: Thyreophora): dermal armour from the Lower Kota Formation (Lower Jurassic) of India. 291, 205–219 (2019).
-
48.
Carpenter, K., Miles, C. & Cloward, K. Skull of a Jurassic ankylosaur. 393, 782–783 (1998).
-
49.
Kirkland, J. I. & Carpenter, K. North America’s first pre-Cretaceous ankylosaur (Dinosauria) from the Upper Jurassic Morrison Formation of western Colorado. 40, 25–41 (1994).
-
50.
Carpenter, K., Miles, C. A., & Cloward, K. in (ed. Carpenter, K.) 55–75 (Indiana Univ. Press, 2001).
-
51.
Galton, P. M. & Carpenter, K. The plated dinosaur Gilmore, 1914 (Dinosauria: Ornithischia; Upper Jurassic, western USA), type species of n. gen. 279, 185–208 (2016).
-
52.
Mateus, O., Dinis, J. & Cunha, P. P. The Lourinhã Formation: the Upper Jurassic to lowermost Cretaceous of the Lusitanian Basin, Portugal – landscapes where dinosaurs walked. 19, 75–97 (2017).
-
53.
Sander, P. M. Longbone histology of the Tendaguru sauropods: implications for growth and biology. 26, 466–488 (2000).
-
54.
Padian, K. & Lamm, E. (Univ. California Press, 2013).
-
55.
Hayashi, S., Carpenter, K. & Suzuki, D. Different growth patterns between the skeleton and osteoderms of (Ornithischia: Thyreophora). 29, 123–131 (2009).
-
56.
Stein, M., Hayashi, S. & Sander, P. M. Long bone histology and growth patterns in ankylosaurs: implications for life history and evolution. 8, e68590 (2013).
-
57.
Waskow, K. & Mateus, O. Dorsal rib histology of dinosaurs and a crocodile from western Portugal: skeletochronological implications on age determination and life history traits. 16, 425–439 (2017).
Delhi’s air quality plunges to ‘severe’ category, Supreme Court suggests lockdown to tackle crisis
A thick blanket of haze enveloped the city on November 13th, as pollution levels soared to the deep end of the ‘severe’ zone – the highest for this season and even worse than the postDiwali spike.
The Supreme Court on Saturday termed the rise in air pollution in DelhiNCR an \”emergency\” situation and asked the Centre and the Delhi government to take emergency measures to improve the air quality. Delhi AirPollution DelhiAirPollution Pollution
Also Readhttps://www.newindianexpress.com/cities/delhi/2021/nov/13/itisanemergencysituationthinkaboutimposinglockdownscondelhipollution2383152.html
Also Readhttps://www.newindianexpress.com/cities/delhi/2021/nov/13/hazethickensdelhiinchestowardsemergencymode2383068.html
For Latest Updates on National News, Clickhttps://www.newindianexpress.com/nation
\r
We are active on social media! Follow us:\r
Facebook: https://facebook.com/thenewindianxpress \r
Twitter: https://twitter.com/NewIndianXpress\r
\r
Stay updated with our Apps: \r
For Android users: http://bit.ly/2EdIaWW\r
iOS users: https://apple.co/2SrVdHf\r
\r
For more videos: http://www.newindianexpress.com/videos\r
For all the important news: http://www.newindianexpress.com/
นอกจากการดูบทความนี้แล้ว คุณยังสามารถดูข้อมูลที่เป็นประโยชน์อื่นๆ อีกมากมายที่เราให้ไว้ที่นี่: ดูเพิ่มเติม

Donald Trump: comments about injecting disinfectant were ‘sarcastic’
Donald Trump insisted on Friday that he was being ‘sarcastic’ when he raised the possibility of disinfectants being injected into people to combat Covid19, prompting the makers of Lysol and Dettol to say its products should ‘under no circumstance’ be used as an internal treatment for the coronavirus. The president instead focused on another potentially dangerous idea: using disinfectant on the hands. Household disinfectant is not intended for use on the body
Subscribe to Guardian News on YouTube ► http://bit.ly/guardianwiressub
Coronavirus US live: Trump tries to backtrack on disinfectant remarks, claiming they were ‘sarcastic’ ► https://www.theguardian.com/world/live/2020/apr/24/coronavirususlivenewstrumpcuomogeorgiacaseslatest
Revealed: leader of group peddling bleach as coronavirus ‘cure’ wrote to Trump this week ► https://www.theguardian.com/world/2020/apr/24/revealedleadergrouppeddlingbleachcurelobbiedtrumpcoronavirus
Support the Guardian ► https://support.theguardian.com/contribute
Today in Focus podcast ► https://www.theguardian.com/news/series/todayinfocus
The Guardian YouTube network:
The Guardian ► http://www.youtube.com/theguardian
Owen Jones talks ► http://bit.ly/subsowenjones
Guardian Football ► http://is.gd/guardianfootball
Guardian Sport ► http://bit.ly/GDNsport
Guardian Culture ► http://is.gd/guardianculture

Study suggests single men prioritize sex less than they did pre-pandemic
A Match.com study claims that 81% of single men say that sex is less important to them now than it was before the pandemic.
The \”Good Day Philadelphia\” team breaks down the latest local, regional and national news, along with information on business, entertainment, sports, weather, traffic and more. It’s everything you need to have a good day!
Subscribe to FOX 29: https://www.youtube.com/fox29philly?sub_confirmation=1
Watch FOX 29 live: https://www.fox29.com/live
More \”Good Day Philadelphia\” on YouTube: https://www.youtube.com/playlist?list=PL5yFQTLO3JXG51DVz1dy7fbAPzzxinZd
Download the FOX 29 News app: https://fox29.onelink.me/dyvt?pid=social\u0026c=youtube\u0026af_web_dp=https%3A%2F%2Fwww.fox29.com%2Fapps
Download the FOX 29 Weather app: https://www.fox29.com/weatherapp
Follow FOX 29 on Facebook: https://www.facebook.com/fox29philadelphia/
Follow FOX 29 on Twitter: https://twitter.com/FOX29philly
Follow FOX 29 on Instagram: https://www.instagram.com/fox29philly
Subscribe to FOX 29’s newsletters: https://www.fox29.com/email
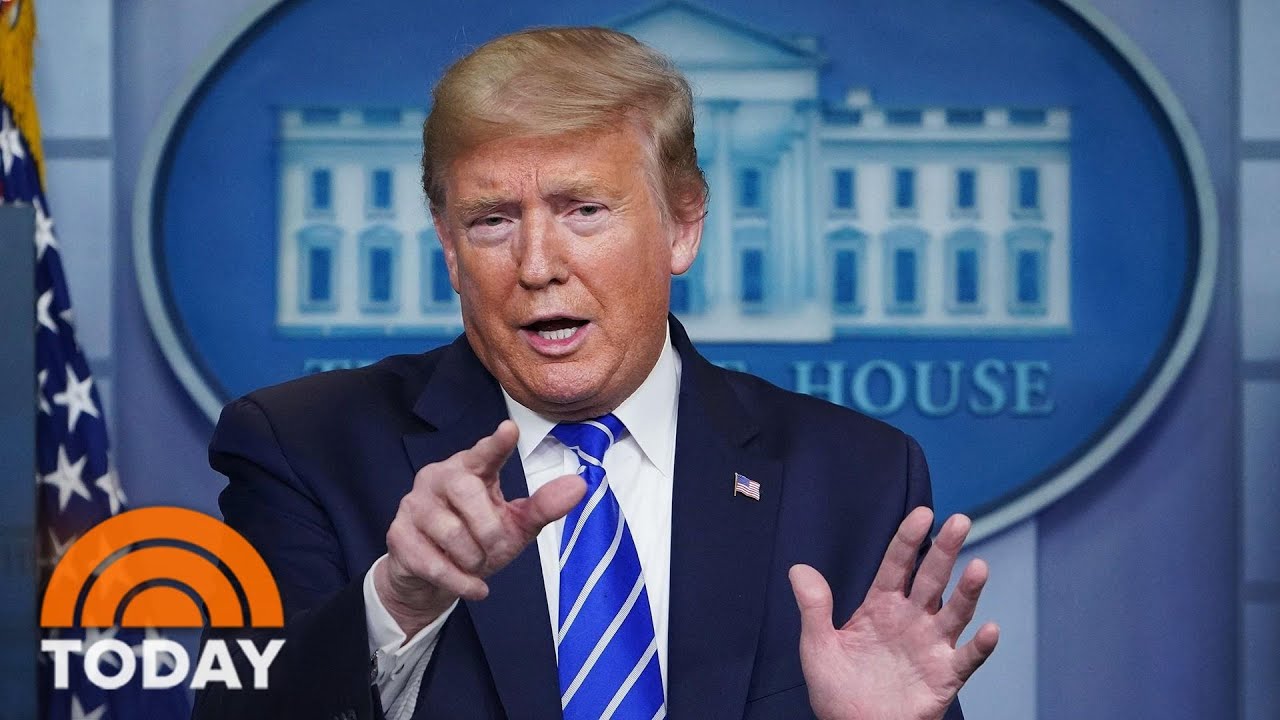
Trump Suggests Light And Disinfectant Treatments For Coronavirus | TODAY
President Trump raised some eyebrows during his daily coronavirus briefing Thursday, suggesting patients be exposed to heat and light or injected with disinfectant. NBC’s Tom Costello reports for TODAY.
» Subscribe to TODAY: http://on.today.com/SubscribeToTODAY
» Watch the latest from TODAY: http://bit.ly/LatestTODAY
About: TODAY brings you the latest headlines and expert tips on money, health and parenting. We wake up every morning to give you and your family all you need to start your day. If it matters to you, it matters to us. We are in the people business. Subscribe to our channel for exclusive TODAY archival footage \u0026 our original web series.
Connect with TODAY Online!
Visit TODAY’s Website: http://on.today.com/ReadTODAY
Find TODAY on Facebook: http://on.today.com/LikeTODAY
Follow TODAY on Twitter: http://on.today.com/FollowTODAY
Follow TODAY on Instagram: http://on.today.com/InstaTODAY
Follow TODAY on Pinterest: http://on.today.com/PinTODAY
Trump Coronavirus TodayShow
Trump Suggests Light And Disinfectant Treatments For Coronavirus | TODAY

Trump floats dangerous coronavirus treatment ideas as Dr Birx looks on
Donald Trump prompted a backlash from medical experts after floating the idea that they could look into heat, light and injections of disinfectants as a cure for Covid19. His public health advisers immediately played down the idea, and medics warn that trying such ideas could be fatal. Coronavirus response coordinator Dr Deborah Birx appeared caught off guard when Trump asked her directly if heat and light would cure the deadly disease. ‘Not as a treatment,’ Birx replied
Bolsonaro won’t help with coronavirus, so Brazil’s favelas are helping themselves ► https://www.youtube.com/watch?v=CbQyUclUSg
Coronavirus US live: Trump says federal distancing guidelines could extend into summer ► https://www.theguardian.com/world/live/2020/apr/23/coronavirususlivehouse484bncrisisaidpackagetrumpcuomolatestnewsupdates
Subscribe to Guardian News on YouTube ► http://bit.ly/guardianwiressub
Support the Guardian ► https://support.theguardian.com/contribute
Today in Focus podcast ► https://www.theguardian.com/news/series/todayinfocus
The Guardian YouTube network:
The Guardian ► http://www.youtube.com/theguardian
Owen Jones talks ► http://bit.ly/subsowenjones
Guardian Football ► http://is.gd/guardianfootball
Guardian Sport ► http://bit.ly/GDNsport
Guardian Culture ► http://is.gd/guardianculture

นอกจากการดูบทความนี้แล้ว คุณยังสามารถดูข้อมูลที่เป็นประโยชน์อื่นๆ อีกมากมายที่เราให้ไว้ที่นี่: ดูวิธีอื่นๆMAKE MONEY ONLINE
ขอบคุณที่รับชมกระทู้ครับ suggests